BASE - Biologie de l’Adaptation et Systèmes en Évolution
17/11/2024 -
GQE
Modélisation de la variation quantitative et de son évolution, en intégrant les niveaux génétique, moléculaire, génomique, métabolique et environnemental.
Head :
Judith Legrand (UPSay)
Head :
Judith Legrand (UPSay)
Our research aims at understanding the dynamics of adaptation and the evolution of life-history traits of populations at different biological scales. The associated projects are structured around three main themes: the genotype-phenotype relationship at different levels of phenotypic integration, the dynamics of adaptation at different time scales and interactions between genotype and the biotic and abiotic environment (GxE). To carry out this research, we use experimental approaches (field, greenhouse and laboratory experiments) as well as mathematical modelling approaches. Although most of our projects focus on maize and yeast, we work on other biological models when they are more appropriate for addressing scientific questions, given the technical constraints and the current state of knowledge.
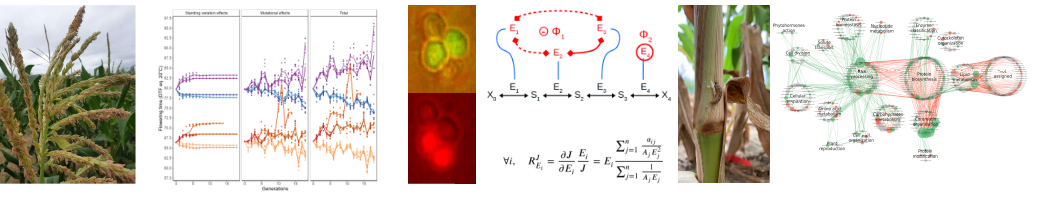
Theme 1. The genotype-phenotype relationship at different levels of phenotypic integration
Understanding the genotype-phenotype (GP) relationship is a central issue in evolutionary biology and plant/animal breeding. The non-linearity of processes shaping this relationship makes it complex.
We are studying this complexity in three projects:
- the basis of heterosis, an emerging property of the complexity of the GP relationship
- the metabolic basis of phenotypic variation in yeast and cultivated plants such as maize
- the evolutionary dynamics of enzyme levels in metabolic pathways. Our approach combines the integration of omics data, mathematical modelling and computer simulations.
Related projects :
- HeteroYeast (ANR) , Metaflux (Plant2Pro)
- Current and past PhD projects (date of defense): Maëla Sémery, Yacine Djabali (2023), Charlotte Coton (2021), Marianyela Petrizzelli (2019)
Theme 2. Dynamics of adaptation
o understand the dynamics of adaptation, we study both short- and long-term evolutionary processes. To address these questions, we use both experimental evolutionary and theoretical approaches. This combination of approaches enables us to study how the different scales of life - from molecular mechanisms to populations - interact to shape the dynamics of adaptation.
Our studies mainly deal with :
- the evolution of the rate of recombination and its role in the dynamics of adaptation
- the evolutionary consequences of sexual versus non-sexual reproductive regimes
- the response to selection on flowering time in maize
- survival at a long stationary phase in Escherichia coli as part of a first-year undergraduate teaching unit.
Related projects :
- Evolrec (ANR) , CO-PATT (ANR) , Itemaize (BASC) , EpiFlore (BAP INRAE), Expérience de Sélection Divergente du Plateau de Saclay.
- Current and past PhD projects (date of defense): Arnaud Desbiez-Piat (2021), Elise Tourrette (2020)
Theme 3: Genotype-environment interactions
In the context of global change, a better understanding of the genetic basis of adaptation and GxE interactions is crucial. We are studying the intraspecific genetic and phenotypic diversity of our biological models (plants, micro-organisms) in relation to environmental variations. The response of phenotypes to environmental variations is observed at different scales, from molecular to integrated traits. We are interested in variations in the abiotic environment, in particular drought, and variations in the biotic environment, in particular pest pressure or the rhizospheric microbial communities.
Our studies mainly deal with :
- the diversity of the response of plants (maize, fruit trees) to climatic variables at different life scales
- the adaptation of species to their environments via the identification of genotype-environment associations
- interactions between plants and their biotic environment (insect pests, weeds, pathogenic fungi, microbial communities in the rhizosphere, crop-predator birds, etc.).
Related projects :
- Itemaize (BASC) , Amaizing (ANR) , WarmRules (DATAIA) , INCREASE (Europe) , PEPR Agroécologie et Numérique , Stat4Plant (ANR) , MYCORE (C-BASC), GAIcirse(C-BASC), PHENOFORE (SEMAE) , BIOCOSMA (ANR), MetaP-Rhiz (BAP, Biosphera)
- Current and past PhD projects (date of defense): Maxime Criado, Antoine Caillebotte, Sacha Revillon (2024), Inoussa Sanane (2020)
Members
- Diala Abu Awad, Assistant professor (UPSay)
- Willy Vincent Bienvenut, Junior Investigator (CNRS), 50% , 50%
- Mélisande Blein-Nicolas, Research Engineer (INRAE), 50% , 50%
- Dominique de Vienne, Professor (UPSay)
- Christine Dillmann, Professor (Université Paris-Saclay, Faculté des Sciences)
- Cécile Fairhead, Professor (UPSay)
- Matthieu Falque, Research Engineer (INRAE), 50% , 50%
- Judith Legrand, Maitre de conférences (UPSay)
- Élodie Marchadier, Assistant Professor (UPSay)
- Xavier Raffoux, Engineer (INRAE), 50% , 50%
- Adrienne Ressayre, Research scientist (INRAE)
- Maëla Sémery, PhD student ()
Publications
2024- Falque M. , Bourgais A., Dumas F., De Carvalho M., Diblasi C.. (2024) MiniRead: A simple and inexpensive do‐it‐yourself device for multiple analyses of micro‐organism growth kinetics. Yeast, yea.3932
- Coton C., Dillmann C. , de Vienne D. . (2023) Evolution of enzyme levels in metabolic pathways: A theoretical approach. Part 2. Journal of Theoretical Biology, (558) 111354
- Desbiez-Piat A., Ressayre A. , Marchadier Élodie. , Noly A., Remoue C., Vitte C., Belcram H., Bourgais A., Galic N., Guilloux ML., Tenaillon MI., Dillmann C. . (2023) Pervasive GxE interactions shape adaptive trajectories and the exploration of the phenotypic space in artificial selection experiments. Genetics, (in press) 2023.01.13.523786
- Desbiez-Piat A., Ressayre A. , Marchadier Élodie. , Noly A., Bourgais A., Galic N., Le Guilloux M., Tenaillon MI., Dillmann C. . (2023) The coupling between mutations effects and environment guides the exploration of phenotypic space as evidenced in artificial selection experiments. DOI.org (Crossref),
- Michel E., Masson E., Bubbendorf S., Lapicque L., Nidelet T., Segond D., Guézenec S., Marlin T., Devillers H., Rué O., Onno B., Legrand J. , Sicard D.. (2023) Artisanal and farmer bread making practices differently shape fungal species community composition in French sourdoughs. Peer Community Journal, (3) e11
- Régnier B., Legrand J. , Calatayud PA., Rebaudo F.. (2023) Developmental Differentiations of Major Maize Stemborers Due to Global Warming in Temperate and Tropical Climates. Insects, 1 (14) 51
- Sanane I., Nicolas SD., Bauland C., Marion-Poll F., Noûs C., Legrand J. , Dillmann C. . (2023) Large genetic variability of maize leaf palatability to european corn borer : metabolic insights. bioRxiv,
- Von Gastrow L., Michel E., Legrand J. , Amelot R., Segond D., Guezenec S., Rué O., Chable V., Goldringer I., Dousset X., Serpolay‐Bessoni E., Taupier‐Letage B., Vindras‐Fouillet C., Onno B., Valence F., Sicard D.. (2023) Microbial community dispersal from wheat grains to sourdoughs: A contribution of participatory research. Molecular Ecology, 10 (32) 2413-2427
- Albert B., Matamoro-Vidal A., Prieu C., Nadot S., Till-Bottraud I., Ressayre A. , Gouyon PH.. (2022) A Review of the Developmental Processes and Selective Pressures Shaping Aperture Pattern in Angiosperms. Plants, 3 (11) 357
- Coton C., Talbot G., Le Louarn M., Dillmann C. , de Vienne D. . (2022) Evolution of enzyme levels in metabolic pathways: A theoretical approach. Part 1. Journal of Theoretical Biology, (538) 111015
- David O., Le Rouzic A., Dillmann C. . (2022) Optimization of sampling designs for pedigrees and association studies. Biometrics, 3 (78) 1056-1066
- de Vienne D. , Capy P.. (2022) Special issue on “The relationship between genotype and phenotype: new insight into an old question”. Genetica, 3-4 (150) 151-151
- Hsu YM., 2022-09, Mining genetic diversity for tomorrow's agriculture, Theses, Université Paris-Saclay
- Hsu YM., Falque M. , Martin OC.. (2022) Quantitative modelling of fine-scale variations in the Arabidopsis thaliana crossover landscape. Quant Plant Bio., (3) e3
- Régnier B., Legrand J. , Rebaudo F., Brent C.. (2022) Modeling Temperature-Dependent Development Rate in Insects and Implications of Experimental Design. Environmental Entomology, 1 (51) 132-144
- de Vienne D. . (2022) What is a phenotype? History and new developments of the concept. Genetica, 3-4 (150) 153-158
- Desbiez-Piat A., Le Rouzic A., Tenaillon MI., Dillmann C. . (2021) Interplay between extreme drift and selection intensities favors the fixation of beneficial mutations in selfing maize populations. Genetics, 2 (219)
- Olvera-Vazquez SG., Alhmedi A., Miñarro M., Shykoff JA., Marchadier Élodie. , Rousselet A., Remoué C., Gardet R., Degrave A., Robert P., Chen X., Porchier J., Giraud T., Vander-Mijnsbrugee K., Raffoux X. , Falque M. , Alins G., Didelot F., Beliën T., Dapena E., Lemarquand A., Cornille A.. (2021) Experimental test for local adaptation of the rosy apple aphid (Dysaphis plantaginea) to its host (Malus domestica) and to its climate in Europe. PCI Ecology, (Pre-registration version)
- Petrizzelli MS., de Vienne D. , Nidelet T., Noûs C., Dillmann C. , Kaleta C.. (2021) Data integration uncovers the metabolic bases of phenotypic variation in yeast. PLoS Comput Biol, 7 (17) e1009157
- Plancade S., Marchadier Élodie. , Huet S., Ressayre A. , Noûs C., Dillmann C. . (2021) A new hypothesis-testing model for phyllochron based on a stochastic process - application to analysis of genetic and environment effects in maize. DOI.org (Crossref),
- Sanané I., Legrand J. , Dillmann C. , Marion-Poll F.. (2021) High-Throughput Feeding Bioassay for Lepidoptera Larvae. J Chem Ecol, 7 (47) 642-652
- Tourrette E., Falque M. , Martin OC.. (2021) Enhancing backcross programs through increased recombination. Genet Sel Evol, 1 (53) 25
- Blein-Nicolas M. , Negro SS., Balliau T., Welcker C., Cabrera-Bosquet L., Nicolas SD., Charcosset A., Zivy M.. (2020) A systems genetics approach reveals environment-dependent associations between SNPs, protein coexpression, and drought-related traits in maize. Genome Res., 11 (30) 1593-1604
- Carrillo-Perdomo E., Vidal A., Kreplak J., Duborjal H., Leveugle M., Duarte J., Desmetz C., Deulvot C., Raffiot B., Marget P., Tayeh N., Pichon JP., Falque M. , Martin OC., Burstin J., Aubert G.. (2020) Development of new genetic resources for faba bean (Vicia faba L.) breeding through the discovery of gene-based SNP markers and the construction of a high-density consensus map. Sci Rep, 1 (10) 6790
- Falque M. , Jebreen K., Paux E., Knaak C., Mezmouk S., Martin OC.. (2020) CNVmap: A Method and Software To Detect and Map Copy Number Variants from Segregation Data. Genetics, 3 (214) 561-576
- Hanemian M., Vasseur F., Marchadier Élodie. , Gilbault E., Bresson J., Gy I., Violle C., Loudet O.. (2020) Natural variation at FLM splicing has pleiotropic effects modulating ecological strategies in Arabidopsis thaliana. Nat Commun, 1 (11) 4140
- Harlé O., Legrand J. , Tesnière C., Pradal M., Mouret JR., Nidelet T., Ohya Y.. (2020) Investigations of the mechanisms of interactions between four non-conventional species with Saccharomyces cerevisiae in oenological conditions. PLoS ONE, 5 (15) e0233285
- Henry V., Saïs F., Inizan O., Marchadier Élodie. , Dibie J., Goelzer A., Fromion V.. (2020) BiPOm: a rule-based ontology to represent and infer molecule knowledge from a biological process-centered viewpoint. BMC Bioinformatics, 1 (21) 327
- Sanane I., Legrand J. , Dillmann C. , Marion-Poll F.. (2020) A semi-automated design for high-throughput Lepidoptera larvae feeding bioassays. bioRxiv, 2020.08.02.232256
- Sanane I., 20 octobre 2020, Composantes de la dynamique de l’interaction entre le maïs et les insectes lépidoptères foreurs de tige, PhD thesis, Université Paris-Saclay
- de Vienne D. , Fiévet JB.. (2020) The Pitfalls of Heterosis Coefficients. Plants, 7 (9) 875
- Bienvenut WV. , 2 décembre 2019, Des débuts de la protéomique à l'acétylation N-terminale des protéines chez les plantes, HDR, Université Paris Saclay
- Jebreen K., Petrizzelli M., Martin OC.. (2019) Probabilities of Multilocus Genotypes in SIB Recombinant Inbred Lines. Front. Genet., (10) 833
- Petrizzelli M., de Vienne D. , Dillmann C. . (2019) Decoupling the Variances of Heterosis and Inbreeding Effects Is Evidenced in Yeast's Life-History and Proteomic Traits. Genetics, 2 (211) 741-756
- Petrizzelli M., 8 juillet 2019, Mathematical modelling and integration of complex biological data: analysis of the heterosis phenomenon in yeas, PhD Thesis, Université Paris-Saclay
- Srisuwan S., Sihachakr D., Martín J., Vallès J., Ressayre A. , Brown SC., Siljak‐Yakovlev S., Arroyo J.. (2019) Change in nuclear DNA content and pollen size with polyploidisation in the sweet potato ( Ipomoea batatas , Convolvulaceae ) complex. Plant Biol J, 2 (21) 237-247
- Termolino P., Falque M. , Aiese Cigliano R., Cremona G., Paparo R., Ederveen A., Martin OC., Consiglio FM., Conicella C.. (2019) Recombination suppression in heterozygotes for a pericentric inversion induces the interchromosomal effect on crossovers in Arabidopsis. Plant J, 6 (100) 1163-1175
- Tourrette E., Bernardo R., Falque M. , Martin OC.. (2019) Assessing by Modeling the Consequences of Increased Recombination in Recurrent Selection of Oryza sativa and Brassica rapa. G3, 12 (9) 4169-4181
- Tourrette E., 25 novembre 2019, Unleashing genetic diversity in breeding by increasing recombination: an in silico study, PhD Thesis, Université de Paris-Diderot
- Urien C., Legrand J. , Montalent P., Casaregola S., Sicard D.. (2019) Fungal Species Diversity in French Bread Sourdoughs Made of Organic Wheat Flour. Front. Microbiol., (10) 201
- Vasseur F., Fouqueau L., de Vienne D. , Nidelet T., Violle C., Weigel D.. (2019) Nonlinear phenotypic variation uncovers the emergence of heterosis in Arabidopsis thaliana. PLoS Biol, 4 (17) e3000214
- Albert B., Ressayre A. , Dillmann C. , Carlson AL., Swanson RJ., Gouyon PH., Dobritsa AA.. (2018) Effect of aperture number on pollen germination, survival and reproductive success in Arabidopsis thaliana. Annals of Botany, 4 (121) 733-740
- Carbonetto B., Ramsayer J., Nidelet T., Legrand J. , Sicard D.. (2018) Bakery yeasts, a new model for studies in ecology and evolution. Yeast, 11 (35) 591-603
- Collot D., Nidelet T., Ramsayer J., Martin OC., Meleard S., Dillmann C. , Sicard D., Legrand J. . (2018) Feedback between environment and traits under selection in a seasonal environment: consequences for experimental evolution. Proceedings. Biological sciences, 1876 (285)
- Fiévet JB., Nidelet T., Dillmann C. , de Vienne D. . (2018) Heterosis Is a Systemic Property Emerging From Non-linear Genotype-Phenotype Relationships: Evidence From in Vitro Genetics and Computer Simulations. Front. Genet., (9)
- Raffoux X. , 2018-06-11 06/11/18, Diversité et déterminisme génétique de la recombinaison méiotique chez Saccharomyces cerevisiae, PhD thesis, Université Paris-Saclay
- Raffoux X. , Bourge M., Dumas F., Martin OC., Falque M. . (2018) High-throughput measurement of recombination rates and genetic interference in Saccharomyces cerevisiae. Yeast, 6 (35) 431-442
- Raffoux X. , Bourge M., Dumas F., Martin OC., Falque M. . (2018) Role of Cis, Trans, and Inbreeding Effects on Meiotic Recombination in Saccharomyces cerevisiae. Genetics, 4 (210) 1213-1226
- Tenaillon MI., Sedikki K., Mollion M., Guilloux ML., Marchadier Élodie. , Ressayre A. , Dillmann C. . (2018) Transcriptomic response to divergent selection for flowering time in maize reveals convergence and key players of the underlying gene regulatory network. bioRxiv, 461947
- Giraud H., Bauland C., Falque M. , Madur D., Combes V., Jamin P., Monteil C., Laborde J., Palaffre C., Gaillard A., Blanchard P., Charcosset A., Moreau L.. (2017) Linkage Analysis and Association Mapping QTL Detection Models for Hybrids Between Multiparental Populations from Two Heterotic Groups: Application to Biomass Production in Maize (Zea mays L.). G3: Genes, Genomes, Genetics, g3.300121.2017
- Henry VJ., Saïs F., Marchadier Élodie. , Dibie J., Goelzer A., Fromion V.. (2017) BiPOm: Biological interlocked Process Ontology for metabolism. How to infer molecule knowledge from biological process?. HAL Archives Ouvertes, np
- Pelé A., Falque M. , Trotoux G., Eber F., Nègre S., Gilet M., Huteau V., Lodé M., Jousseaume T., Dechaumet S., Morice J., Poncet C., Coriton O., Martin OC., Rousseau-Gueutin M., Chèvre AM.. (2017) Amplifying recombination genome-wide and reshaping crossover landscapes in Brassicas. PLoS Genet, 5 (13)
- Sabarly V., Aubron C., Glodt J., Balliau T., Langella O., Chevret D., Rigal O., Bourgais A., Picard B., de Vienne D. , Denamur E., Bouvet O., Dillmann C. . (2016) Interactions between genotype and environment drive the metabolic phenotype within Escherichia coli isolates. Environmental microbiology, 1 (18) 100-17